
Cialis ist bekannt für seine lange Wirkdauer von bis zu 36 Stunden. Dadurch unterscheidet es sich deutlich von Viagra. Viele Schweizer vergleichen daher Preise und schauen nach Angeboten unter dem Begriff cialis generika schweiz, da Generika erschwinglicher sind.
Ibmb.csic.es
The antimalarial and
people suffering from the disease per year. Africa accounts forover 90% of reported cases, with an annual 20% increase of
cytotoxic drug cryptolepine malaria-related illness and death. Malaria is responsible for as
many deaths per annum as AIDS for all of the last 15 years. Drug
intercalates into DNA at
resistance to malaria has become one of the most significantthreats to human health and the search for new effective drugs is
urgent. Although the mechanism of action of the antimalarialdrugs is unclear, many of these drugs, such as chloroquine and
John N. Lisgarten1,2, Miquel Coll1, Jose Portugal1,
quinacrine, are known to interact with DNA1.
Colin W. Wright3 and Juan Aymami1,4
Cryptolepine (5-methyl indolo[2,3b]-quinoline) is an indolo-
quinoline alkaloid first isolated from the roots of Cryptolepsis
1Institut de Biologia Molecular de Barcelona, C.S.I.C., Jordi Girona 18,
triangularis collected in Kisantu (Congo). Extracts of the roots
08034 Barcelona, Spain. 2Department of Crystallography, Birkbeck College,
of the related climbing liana Cryptolepsis sanguinolenta, in which
University of London, Malet Street, London, WC1E 7HX, UK. 3The School of
cryptolepine is the main alkaloid, have been used clinically in
Pharmacy, University of Bradford, West Yorkshire, BD7 4ER, UK.
Ghana for the treatment of malaria2, and also as a remedy
4Department d'Enginyeria Quimica, Universitat Politècnica de Catalunya,
against colic and stomach ulcers. Cryptolepine itself has been
Diagonal 647, 08028 Barcelona, Spain.
found to produce a variety of pharmacological effects, includinghypotensive and antipyretic properties, presynaptic α-adreno-
Published online: 3 December 2001, DOI: 10.1038/nsb729
receptor blocking action, antimuscarinic properties, anti-inflammatory properties and antibacterial effects (for review
Cryptolepine, a naturally occurring indoloquinoline alka-
see ref. 3).
loid used as an antimalarial drug in Central and Western
Cryptolepine has potent in vitro activity against the malaria
Africa, has been found to bind to DNA in a formerly
parasite (Plasmodium falciparum) and possesses cytotoxic activ-
unknown intercalation mode. Evidence from competition
ity, inhibiting DNA synthesis in B16 melanoma cells3. The alka-
dialysis assays demonstrates that cryptolepine is able to bind
lishing Gr
loid was found to bind tightly to DNA and behaved as a typical
CG-rich sequences containing nonalternating CC sites. Here
intercalating agent. The drug interacts preferentially with CG-
we show that cryptolepine interacts with the CC sites of the
rich sequences and discriminates against homooligomeric runs
DNA fragment d(CCTAGG)2 in a base-stacking intercalation of A and T. The study3 also led to the discovery that cryptolepine
mode. This is the first DNA intercalator complex, from ∼90 is a potent topoisomerase II inhibitor and a promising anti-
solved by X-ray crystallography, to bind a nonalternating
tumor agent. Cryptolepine stabilizes topoisomerase II–DNA
(pyrimidine-pyrimidine) DNA sequence. The asymmetry of
covalent complexes and stimulates the cutting of DNA at a sub-
2002 Nature Pub
the drug induces a perfect stacking with the asymmetric site,
set of preexisting topoisomerase II cleavage sites3,4. In addition,
allowing for the stability of the complex in the absence of
evidence suggests that cryptolepine may inhibit the detoxifica-
hydrogen bonding interactions. The crystal structure of this
tion of heme produced by malaria parasites in red blood cells as
antimalarial drug–DNA complex provides evidence for the
a result of the digestion of hemoglobin, similarly to chloroquine
first nonalternating intercalation and, as such, provides a
and related 4-aminoquinoline antimalarials5. Although the anti-
basis for the design of new anticancer or antimalarial drugs.
malarial activity of cryptolepine may involve a chloroquine-like
Malaria, by far the most important tropical parasite, causes an
action, interactions with DNA may also contribute. This is sup-
estimated annual 2.7 million deaths among the 300–500 million
ported by a fluorescence microscopy study, which suggests that
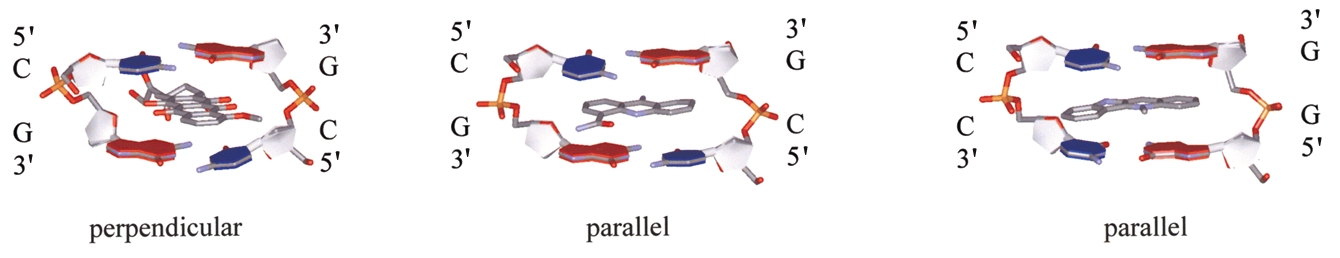
Fig. 1 Diagram showing the main intercalation modes. The cryptolepine site does not have two-fold symmetry. Coordinates are taken from the
Nucleic Acid Data Base for a, the anthracycline type d(CGCGCG)–epidoxorubicin (NDB code dd0022); b, the acridine type d(CGTACG)–9-amino-DACA
(NDB code dd0015); and c, d(CCTAGG)–cryptolepine (this paper).
nature structural biology • volume 9 number 1 • january 2002
Fig. 2 Results obtained from the Ren and Chaires competition dialysis
experiment for cryptolepine. The amount of ligand bound to each DNA
fragment is shown graphically. The sites provided for each fragment are
shown in columns.
tolepine interacts with the site d(CpC)-d(GpG) in a base-stack-ing intercalation mode.
Cryptolepine binding to DNA in solution
The Ren and Chaires competition dialysis method20 was used todetermine the sequence selectivity of cryptolepine for differentsmall DNA fragments. In order to determine the preference ofthe ligand for base sequences of alternating and nonalternatingC-G (CC and CG, respectively), and alternating and nonalter-
cryptolepine accumulates into parasite structures that may cor-
nating A-T (AA and AT, respectively) base pairs, fragments of
respond to the parasite nucleus6. Cryptolepine was also localized
the same size were selected. This ensured that the same number
into the kinetoplast DNA of the trypanosome. Curiously, the
of possible similar sites were available for ease of comparability.
DNA sequences in the minicircles of kinetoplast contain a larger
The competition dialysis assay results indicated that cryp-
number of CC/GG sites (∼3:1) than CG/GC (http://www. tolepine prefers to bind to C-G rich sequences of DNA, with aebi.ac.uk:80/parasites/kDNA/Source.html). The intercalating
tendency for nonalternating CC (Fig. 2). The small differences
properties of cryptolepine were deemed worthy of investigation
between AA and AT are probably not significant. Comparison of
because this compound may lead to new antiprotozoal and anti-
the binding affinity of cryptolepine with those of the ligands
cancer drugs.
used in the original Ren and Chaires experiment indicates thatthe binding affinity of cryptolepine is similar to that of other
lishing Gr
intercalators, such as actinomycin D, daunomycin, porphyrin
Two main kinds of noncovalent DNA–drug interactions are
compounds and chromomycin20. Previous footprinting analy-
known: base intercalation7 and minor groove binding8. Minor
ses3,4 showed that tracts containing CC-GG, as well as several
groove binders that specifically target DNA have been thorough-
CG-GC sites were protected from DNase I cleavage, which is
ly investigated by Dervan9 using hairpin iminopyridines, which
consistent with our binding data (Fig. 2).
allow a proper recognition of DNA sequences. Intercalators arethe group of compounds that bind between the bases of DNA,
2002 Nature Pub
thereby interrupting transcription, replication and/or topoiso-
The complex d(CCTAGG)–cryptolepine has been crystallized,
merase activities10. Although some intercalators have been used
and its structure solved and refined to 1.4 Å resolution (Table 1;
as anticancer drugs, others are carcinogens. Bisintercalators11,
Fig. 2c). The main feature of this structure is the perfect fit of the
trisintercalators12, tetraintercalators13 and octakis-intercala-
drug sandwiched between two consecutive C-G base pairs form-
tors14, containing the same repeating intercalator group, have
ing the first nonalternating site (CC)-(GG)–cryptolepine
been synthesized. Cryptolepine is the first intercalator that
(Figs 1,3). The aromatic six-membered ring of the cryptolepine
appears to prefer or tolerate nonalternating steps (pyrimidine-
molecule stacks between the two cytosines, whereas the fused
aromatic, double six-membered ring portion of the molecule
Two main kinds of DNA–drug intercalation are observed. The
stacks between two guanines (Fig. 3d). The five-membered ring,
first is perpendicular intercalation, typified by doxorubicin and
placed in the middle, gives asymmetry to the cryptolepine mole-
daunomycin7, in which long fused-ring molecules penetrate per-
cule and separates both aromatic groups. The positively charged
pendicularly to the base pair hydrogen bonds. Parallel base-
N16 atom (quinoline group) between the two O6 atoms of con-
stacking intercalators, such as actinomycin15 and acridine-type
secutive guanines in the major groove of DNA and the N8
drugs16, which intercalate parallel to the base pair hydrogen
(indole nitrogen) between the O2 atoms of adjacent cytosines in
bonds and stack their aromatic rings into the DNA bases (Fig. 1),
the minor groove both enhance the stability of the complex
exemplify the other type of intercalation. Perpendicular interca-
(Fig. 3c). This positively charged N16 nitrogen placed in the
lators mainly go to CG and other alternating pyrimidine-purine
major groove between oxygens is also observed in the structure
sites, such as TG or CA. Parallel base-stacking intercalators can
of the complex of 9-amino-DACA interacting with d(CGTACG)
also go to nonalternating sequences. At present there are ∼90 in the CG–drug site16. In this case, the charged nitrogen is placedstructures of nucleic acid–intercalator complexes in the Nucleic
between two oxygens from guanines in different strands; howev-
Acid Database17 that have been shown to have alternating-base
er, the charged nitrogen in the present structure is placed
intercalation sites, most being CG, a few GC and some TG.
between oxygens of adjacent guanines of the same strand. The
The NMR structure of two intercalators, esperamicin A 18
cryptolepine molecule is slightly bent, with a 6.8° angle between
calicheamicin γ 19
1 , in complex with DNA indicate that they
the two aromatic rings, which is similar to the 4.8° angle found
intercalate a single aromatic ring at a CC site. However, the small
in the high resolution X-ray structure of the cryptolepine
size of the intercalator and its two minor groove binding groups
tetraphenyl borate21. The presence of the five-membered ring
suggest that the sequence specificity of the drug is favored by the
positioned between the two aromatic groups allows this bend.
complementarity of the fit between the drug and the floor of the
Neocryptolepine, an isomer of cryptolepine found to a lesser
minor groove. These are minor groove binders, placing a six-
extent than cryptolepine in the plant extracts, shows a reduced
member aromatic ring between CC. The specificity is through
affinity for DNA22. In this isomer, the charged group
the minor groove and not through the intercalation. The crystal
N16-C18H3 of the quinoline moiety is interchanged with the C6
structure reported here shows, for the first time, how cryp-
from the indole (Fig. 1), with both nitrogens on the same side of
nature structural biology • volume 9 number 1 • january 2002

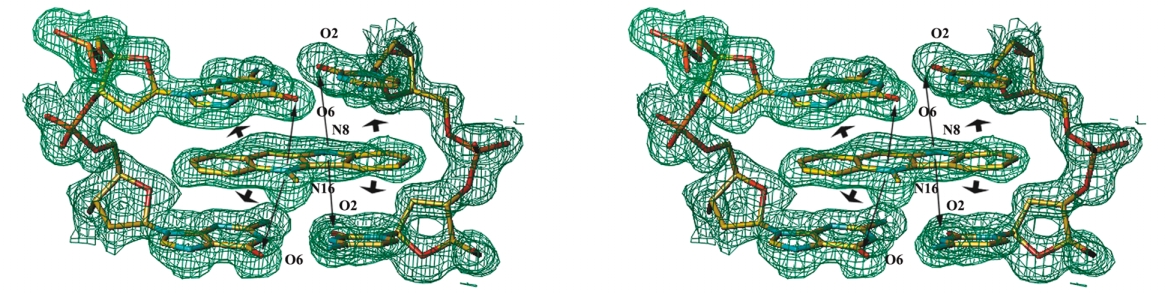
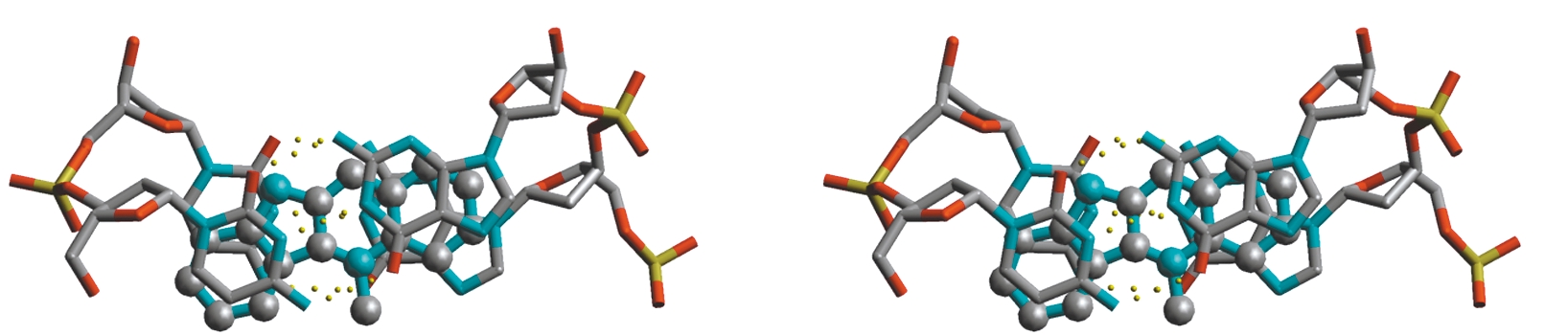
Fig. 3 Crystal structure of the complex. a
a, Scheme for the DNA–cryptolepine com-
plex. b, Stereo view of two bis-intercalat-
ed d(CCTAGG)2 hexanucleotides in the
ab-plane, with the end-stacked ligand
bound between them. Four asymmetric
units are represented in different colors.
c, Stereo view of the 2Fo – Fc electron den-
sity map at the area of the intercalated
ligand, looking into the major groove.
The map was contoured at the 1.2 σ level.
Stacking (large arrows) and electroctatic
(small arrows) interactions are shown.
d, Stereo view of the projection down the
helix axis of a d(CpC)-d(GpG) dinucleotide
with the sandwiched ligand.
the molecule. This reduced DNAaffinity can be understood in termsof the reduced stability of the mole-cule within the complex. In the
neocryptolepine molecule, the per-fect fit that exists in the cryptolepine
complex on both sides of the DNA inthe major and minor grooves, wherethe nitrogen atoms are placedbetween oxygens (Fig. 3c), is impos-
lishing Gr
sible because both nitrogen atomsare on the same side. The cryp-
tolepine molecule has no hydrogen-bonding contacts either with basesof the hexanucleotide or with sol-vent. The absence of such interac-tions suggests that stacking forces
2002 Nature Pub
alone provide the stabilizing mecha-
nism of the complex. The stackinginteractions between the intercalated ligand and the DNA bases
across the stacked ligand (Fig. 3b). These DNA columns are per-
(Fig. 3d) show that the cryptolepine is aligned with its major axis
pendicular to the c-axis and rotated with respect to the neigh-
parallel to the Watson and Crick hydrogen bonds of the base
bors, introducing the phosphate backbone in the minor groove
pairs. The positively charged cryptolepine chromophore is near-
of the neighboring DNA column.
ly enveloped by the two base pairs at the intercalation site andpenetrates deeply into the helical stack, forming strong
hydrophobic interactions with the base pairs and positioning its
The DNA in the complex has a B-like conformation, with
center of mass as close to the helix axis as possible, where the
Watson-Crick base pairing. However, in accommodating the
negative electrostatic potential of the DNA is the greatest23. In
intercalated cryptolepine molecule, the DNA assumes confor-
this way, the chromophore comes to lie in a position where both
mational parameters significantly different from average B-DNA
its hydrophobic and electrostatic interactions are maximized.
values. Nevertheless, the DNA structure is similar to other DNA
The analysis of solved DNA–drug complexes reveals the
complexes with base stacking intercalators, such as proflavine25
importance of stacking forces. Calorimetric and spectroscopic
and 9-amino-DACA16. In the intercalation cavity, the bases are
studies of the compound Hoechst 33258 in complex with
separated by 7 Å, which is much larger than the 5–6 Å observed
d(CGCAAATTTGCG)2 shows that hydrogen bonds contribute in the case of anthracycline drugs daunomycin and doxorubicin.
little to the stability of the complex compared to hydrophobic
In these cases, the drug intercalates perpendicular to the hydro-
gen bonds of the base pairs, causing a buckle on base pairs of theDNA site (Fig. 1). In the present case, the major axis of the drug
aligns parallel to the major axis of the base pairs to maximally
The asymmetric unit contains one strand of DNA hexamer, one
occupy the intercalation site. To achieve this major opening of
intercalated cryptolepine molecule, 37 ordered water molecules
the bases, the sugar-phosphate backbone makes a coupled rota-
and an additional cryptolepine molecule located on the two-fold
tion of the α/γ main torsion angles at cytosine C2, bringing the
axis, sandwiched between the two DNA hexamers. The crystallo-
oxygen O2P into the major groove. On the opposite chain, the
graphic two-fold axis is coincident with the large axis of the drug
same opening is achieved at guanine G6 by small variations in all
molecule. This additional drug molecule links contiguous DNA
the torsion angles. The DNA puckering is generally C2′-endo,
hexamers in the crystal to form a continuous column of duplex-
except for the first cytidine, which is C3′-endo. At the intercala-
es. Because the end-stacked ligand lies on a two-fold axis coinci-
tion sites, the sugars of cytidine C1 and C2 adopt the conforma-
dent with its major axis, the polarity of the DNA backbone
tion C3′-endo and C2′-endo, a configuration frequently found in
reverses at this point, bringing the 5′ termini of adjacent helices intercalator–dinucleotide monophosphate complexes16. DNAinto close juxtaposition, and the cytosine C1 oppose each other
helical twist at the intercalation site is 24°, being unwound by 12°
nature structural biology • volume 9 number 1 • january 2002
Crystal structure resolution. Crystals were grown by mixing
Table 1 Crystallization data and refinement statistics
0.5 µl of 5 mM cryptolepine hydrochloride and 0.5 µl of 3 mM
d(CCTAGG) with 1.0 µl of the crystalization solution containing
Unit cell dimensions
5 mM magnesium acetate, 25 mM 2-(N-morpholino)propanesulfon-
ic acid (MES), pH 6.5, and 1.25 M ammonium sulfate. Single crystalswere flash-frozen in a stream of evaporating liquid nitrogen at
120 K. Diffraction data were collected at EMBL beamline BW7A
(DESY, Hamburg). The structure was solved by molecular replace-
ment using DNA coordinates of the d(CGTACG)–9-amino-DACA
structure, without the drug as starting model, with AMoRe27.
Refinement followed with CNS28, first as a rigid body. The optimum
Number of unique data
orientation of the intercalated cryptolepine was identified by plac-ing the drug at each of the four possible positions, and refining
Completeness1 (%) 87.3
until the best fit and corresponding best R-factor (28.2%) and R
(33.2%) were found. At this stage, an iterative refinement proce-
<I / σ (I)>1
dure was carried out using SHELX-97 (ref. 29), interspersed with
inspection of electron density maps, water positioning and manual
model rebuilding with TURBO-FRODO30. For cryptolepine, bond
lengths and bond angles were refined to specified target values
obtained from the cryptolepine structure determined by Wright
R.m.s. deviation from ideality
et al.21 The DNA bases and the two fused ring system of the cryp-
Bond length (Å)
tolepine were restrained to planar, whereas all other torsion angles
Bond angle distances (Å)
remained unrestrained. No hydrogen bond restraints were used
Coordinates. The coordinates have been deposited in the Protein
Data Bank (accession code 1K9G).
1Number in parenthesis is for the last shell (1.45–1.40 Å).
2R
i – Ι/ ∑h
where (Ι) is the mean intensity of reflection
lishing Gr
Rfree is for 5% of total reflections.
This work was supported by grants from the Ministerio de Educacion y
Determined without solvent.
Cultura of Spain and the Generalitat de Catalunya. J.N.L. acknowledges supportfrom MEyC of Spain. Synchrotron data collection was supported by the ESRF and
with respect to standard B-DNA. The adjacent CpT step is also
EU grants to the EMBL-DESY.
unwound with a twist of 27°, whereas the central TpA step isoverwound, with a twist of 51°. This is a very large twist for Correspondence should be addressed to J.A. email: [email protected]
2002 Nature Pub
B-DNA and leaves the base pairs with minimal overlap of their
aromatic rings.
Received 14 June, 2001; accepted 24 October, 2001.
1. Rivas, L., Murza, A., Sanchez-Cortes, S. & Garcia-Ramos, J.V. J. Biomol. Struct.
Dyn. 18, 371–383 (2000).
2. Boye, G.I. & Ampofo, O. Proceedings of the first international seminar on
The perpendicular intercalators (daunomycin and doxorubicin)
cryptolepine (eds Boakye-Yiadom, K. & Bamgbose S.O.A.) 37 (University of
prefer alternating sites (CG), which allow them to be located at
Kumasi, Ghana; 1983).
3. Bonjean, K. et al. Biochemistry 37, 5136–5146 (1998).
the center of mass of the DNA. In contrast, the parallel base-
4. Dassonneville, L. et al. Biochemistry 38, 7719–7726 (1999).
stacking drugs may adapt more readily to the nonalternating
5. Wright, C.W. et al. J. Med. Chem. 44, 3187–3194 (2001).
6. Arzel, E. et al. J. Med. Chem. 44, 949–960 (2001).
sites (CC), leading to better stacking interaction. In addition, the
7. Frederick, C.A. et al. Biochemistry 29, 2538–2549 (1990).
asymmetry of the drug enhances its fitting with the target in the
8. Aymami, J., Nunn, C.M. & Neidle, S. Nucleic Acids Res. 27, 2691–2698 (1999).
9. White, S., Szewczyk, J.W., Turner, J.M., David, E.E. & Dervan, P.B. Nature 391,
case of nonalternating sites (CC)-(GG) because it allows a differ-
468–471 (1998).
ent stacking on one side (CC) or the other (GG) (Fig. 1).
10. Portugal, J. et al. Curr. Med. Chem. 8, 1–8 (2001).
11. Hu, G.G. et al. Biochemistry 36, 5940–5946 (1997).
12. Takenaka, S., Nishira, S., Kondo, H. & Takagi, M. Nucleic Acid Symp. Ser. 27,
71–72 (1992).
13. Lokey, R.S. et al. J. Am. Chem. Soc. 119,7202–7210 (1997).
Ren and Chaires competition dialysis experiment for cryp-
14. Murr, M.M. et al. Bioorg. Med. Chem. 9, 1141–1148 (2001).
tolepine. Different DNA fragments were dialyzed against a com-
15. Shinomiya, M., Chu, W., Carlson, R.G., Weaver, R.F. & Takusagawa, F. Biochemistry
mon ligand solution, and their total concentration within each
34, 8481–8491 (1995).
16. Adams, A., Guss, J.M., Collyer, C.A., Denny, W.A. & Wakelin, L.P.G. Biochemistry
dialysis bag was determined spectrophotometrically. The extinction
38, 9221–9233 (1999).
coefficients for the DNA fragments used in the experiment were
17. Berman, H.M. et al. Biophys. J. 63, 751 (1992).
obtained by the Boser approach26 and was found to be
18. Kumar, R.A., Ikemoto, N. & Patel. D.J. J. Mol. Biol. 265, 173–186 (1997)
19. Kumar, R.A., Ikemoto, N. & Patel. D.J. J. Mol. Biol. 265, 187–199 (1997)
28,600 M–1 cm–1 at 369 nm for cryptolepine3. In the assay, the same
20. Ren, J. & Chaires, J.B. Biochemistry 38, 16067–16075 (1999).
procedures and solutions were used as described in the original
21. Wright, C.W., Phillipson, J.D., Lisgarten, J.N. & Palmer, R.A. J. Chem. Crystallogr.
experiment20. A buffer consisting of 6 mM Na
29, 449–455 (1999).
2HPO4, 2 mM NaH2PO4,
1 mM NaEDTA and 185 mM NaCl, pH 7.0, was used. In this buffer, all
22. Bailly, C. et al. Anticancer Drug Des. 15, 191–201 (2000).
23. Dean, P.M. & Wakelin, L.P.G. Proc. R. Soc. Lond. B 209, 453–471 (1980).
DNA fragments, including d(CGCGCGCG), should be in the B-DNA
24. Haq, I., Ladbury, J.E., Chowdhry, B.Z., Jenkins, T.C. & Chaires, J.B. J. Mol. Biol. 271,
form. In the experiment, 0.5 ml Spectro/Por DispoDialyzer units with
244–257 (1997).
membrane pores of 2,000 Da were used. A second experiment using
25. Shieh, H.-S., Berman, H.M., Dabrow, M. & Neidle, S. Nucleic Acids Res. 8, 85–97
a membrane pore size of 1,000 Da was performed with similar
26. Boser, P.N. Handbook of biochemistry and molecular biology 3rd edn, Vol 1
results. The values shown are for both experiments (mean ± stan-
Nucleic acids (ed. Fasman, B.G.D.) 589 (CRC Press, Boca Raton, Florida; 1976)
dard deviation). During the experiment, all the DNA fragments are
27. Navaza, J. Acta Crystallogr. A 50, 157 (1994).
in equilibrium with the same free ligand concentration so that the
28. Brünger, A.T. et al. Acta Crystallogr. D 54, 905 (1998).
29. Sheldrick, G.M. The SHELX-97 manual (University of Gottingen, Germany; 1997).
amount of cryptolepine bound to each fragment is proportional to
30. Roussel, A. & Campillau, C. Turbo-Frodo. In Silicon graphics geometry partners
the association constant for ligand binding.
directory. 77–79 (Silicon Graphics, Mountain View, California; 1989).
nature structural biology • volume 9 number 1 • january 2002
Source: http://www.ibmb.csic.es/filesusers/Cryptolepine.pdf
VOL. 1 N0 3. JAN -JUNE, 2004 An interview with the D.G -Nigerian Institute of Me dical Research By NIMRNEWS Editorial Board NIMRNEWS: Sir, what will you death. Talking about consider as your achievement in the past subvention, for the This edition of NIMRNews focused more on facts finding mission by engaging the DIRECTOR-GENERAL: The first thing
Magnesium Research 2010; 23 (2): 1-13 Magnesium and cardiovascular system Leviev Heart Center, Chaim Sheba Medical Center, Tel Hashomern and the Sackler Facultyof Medicine, Tel Aviv University, Ramat Aviv, IsraelCorrespondence: M.Shechter, MD, MA, FESC, FACC, FAHA, FACN, Director, Clinical Research Unit, Leviev Heart Center, Chaim Sheba Medical Center, 52621 Tel Hashomer, Israel








